Setup
| Parameter | Value |
|---|---|
| Dimensions | 3D (torsion, rCC, z_lift) |
| Method | SA-CASSCF(2,2) |
| Seeds | 42, 137, 2011 |
| Steps | 15 |
| Population | 12 → 30 |
Score variant legacy. Main tables: regional batch validation_regional_3seeds_20260403_155239. Additional full regional run (seed 42 only): validation_20260403_211825_legacy_nd3_regional_s42 — its zor_seed_42/probes.jsonl is byte-identical to seed 42 in the 3-seed batch (cmp exit 0).
What this demonstrates
The evidence here is not that inZOR-ND identifies a single privileged “right” region in internal coordinates. It is that, in 3D, the run visits and ranks several separated low-gap regions (clusters, sector weights, pooled scatter) — i.e. multi-basin quasi-degeneracy, not funnel collapse to one geometry.
Best gap (smallest S₀–S₁ seen in a run) is one scalar summary per seed. Landscape structure — where probes pile up, how many DBSCAN basins appear, whether the argmin sits in a supported patch — is a separate object: you need both, but the global minimum alone does not describe the geometry-space organisation this page is about.
Main results (per seed)
| Seed | Best gap (eV) | First <10 meV (probe #) | Top-K dominant region (K=50) |
|---|---|---|---|
| 42 | ~2.6×10−4 | 4 | B (~260°; circular mean ~238°) |
| 137 | ~2.0×10−4 | 87 | A (~70–90°) |
| 2011 | ~7.3×10−4 | 316 | B (~240–260°) |
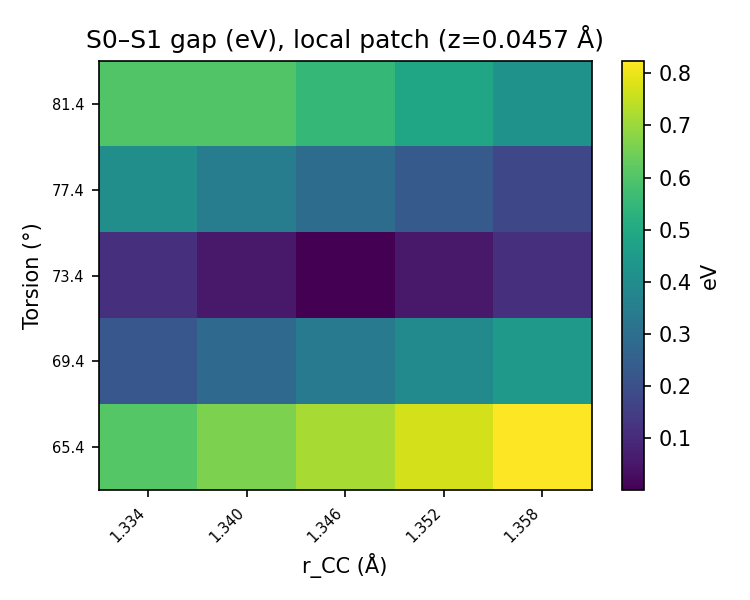
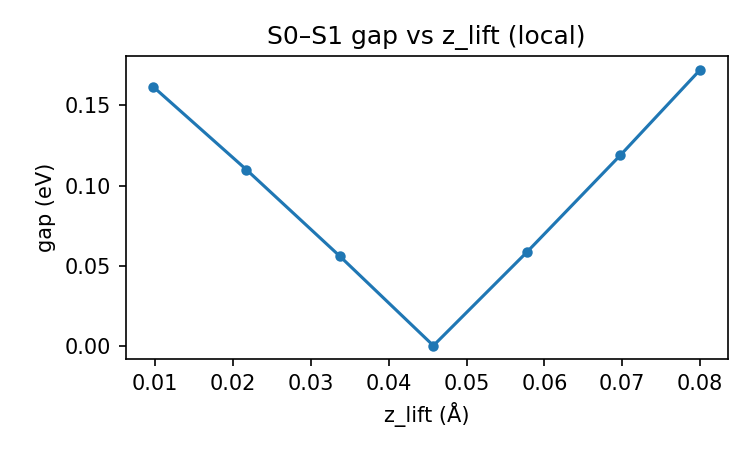
validation_regional_3seeds_summary.json, best_appears_isolated is false for seeds 42, 137, and 2011: the minimum-gap geometry has real local support in the patch metric. The local 5×5 grid and zlift sweep figures below visualize that coherence around the best point.
inZOR-ND vs uniform random (matched budget)
| Metric | inZOR-ND | Random |
|---|---|---|
| Best gap | ~2.0×10−4 eV (best over seeds) | ~5.7×10−4 eV |
| First <10 meV | 4 (fastest seed: 42) | 141 |
Control: n_eval = 8768 uniform samples in [0,1]3 (same internal-coordinate mapping as this 3D engine). inZOR-ND reaches a sub-10 meV gap far earlier than uniform sampling on this budget (e.g. first hit at probe 4 vs 141 in this batch).
Figures (this 3D report’s validation run)
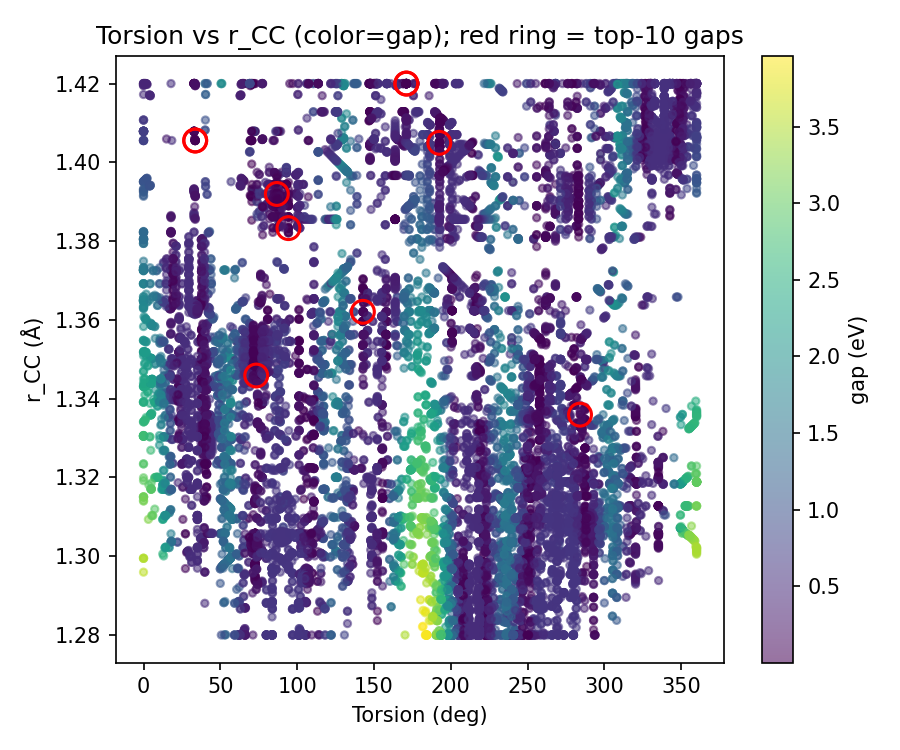
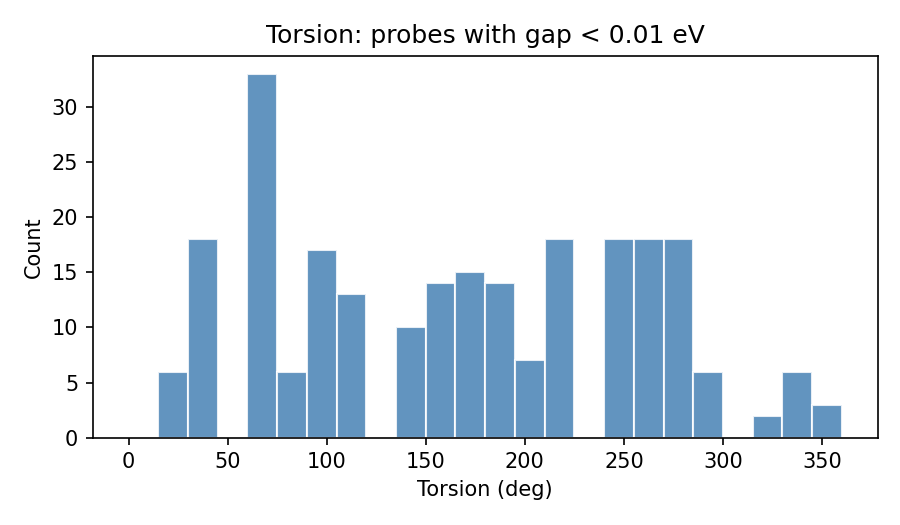
These PNGs are emitted by validate_ethylene_zor.py into logs/validation_regional_3seeds_20260403_155239/ (3 seeds, 3D regional mode) — not ad-hoc graphics from unrelated folders.
Pooled vs per-seed: the scatter, low-gap histogram, local grid, and z sweep are built from concatenated probes across seeds (same logic as regional sections H–J in the full validation write-up). Per-seed regional metrics (sector weights, first <10 meV, isolation flags, etc.) live in validation_regional_3seeds_summary.json — summarized in the main table above, not re-plotted here line-by-line.
Cluster analysis (gap < 0.01 eV, all seeds pooled)
DBSCAN on (cos θ, sin θ, rCC, zlift); hyperparameters match the regional validation cluster export used for this report.
- Total low-gap points: 242
- Total clusters: 15
- Largest cluster: ~11% of non-noise points (16 / 149)
- Substantial clusters (rule in JSON): 6
- Multi-seed clusters: 6
| Cluster | Torsion (°) | Size | Multi-seed |
|---|---|---|---|
| C1 | ~70 | 14 | yes |
| C2 | ~171 | 15 | yes |
| C3 | ~258 | 12 | yes |
| C4 | ~91 | 16 | no |
| C5 | ~252 | 9 | yes |
| C6 | ~145 | 8 | no |
Rows ordered by scientific interest; sizes and angles from exported centroids.
Pooled vs per-seed: Cluster analysis is performed on pooled probes across all seeds, while regional consistency metrics are evaluated per seed.
Interpretation
Regional consistency
- 2 of 3 seeds put more Top-K weight in sector B than A; the third seed (137) concentrates differently — real heterogeneity between runs.
- Concrete Top-K=50 sector-B fractions from the regional summary: ≈0.42 (seed 42), ≈0.08 (seed 137), ≈0.30 (seed 2011).
- In all seeds, the best-gap configuration is not isolated and has local support in the surrounding region.
- All seeds:
best_appears_isolated: false— the best-gap geometry sits in a supported local patch (see figures + JSON). - Low-gap and top-K distributions differ by seed, but concentration in chemically plausible torsion bands persists.
Reproducibility & cluster stability (latest technical step)
If you skim only one line: this section shows determinism when the seed and protocol are fixed (rerun → same probes.jsonl). It is not evidence that the algorithm would match itself across different seeds or across two independent trajectories that actually diverge at the file level.
This does not imply stability across independent runs that generate different probe trajectories. The HIGH verdicts in the table apply only when
cmp on probes.jsonl is identical (byte-for-byte).
Scripts (under scene_ethylene_quasidegen/ in the source tree): low_gap_cluster_analysis.py — cluster_probes_jsonl(...) for one probes.jsonl (no multi-seed aggregation). compare_low_gap_cluster_stability.py — compares run A vs B (--dir-a / --dir-b or explicit paths), re-clusters low-gap points with the same DBSCAN feature space (cos/sin torsion, rCC, zlift), scales on the union of low-gap points, then reports match rate, centroid distances, approximate overlap, and verdict high / moderate / low.
| Comparison (seed 42) | cmp on probes.jsonl | Low-gap (<0.01 eV) | Clusters A/B | Matched | Mean centroid Δ (scaled) | Verdict |
|---|---|---|---|---|---|---|
validation_20260403_130511…_regional_s42 vs …155239/zor_seed_42 | identical (exit 0) | 92 / 92 | 7 / 7 | 7/7 | 0 | HIGH |
…155239/zor_seed_42 vs validation_20260403_211825…_regional_s42 | identical (exit 0) | 92 / 92 | 7 / 7 | 7/7 | 0 | HIGH |
Machine-readable output for the second row is mirrored here: ethylene_3d_artifacts/cluster_stability_rerun_211825_vs_155239.json and ethylene_3d_artifacts/CLUSTER_STABILITY_RERUN_211825_vs_155239.txt.
Same-run reproducibility: identical
probes.jsonl → identical low-gap DBSCAN clusters here (HIGH, zero centroid drift) — validates determinism + clustering code path for that trace only.Still open: cluster stability for two independent runs that produce different probe streams under the same protocol.
Limitations
- Global best-gap geometry is not guaranteed to fall inside fixed A/B sectors.
- Dominant top-K region varies between seeds.
- Clusters are multiple basins, not a single basin.
Conclusion
This result should not be interpreted as convergence to a single optimal geometry.
Instead, the system exhibits multiple quasi-degenerate regions in geometry space. inZOR-ND exploration does not select a unique minimum, but efficiently discovers several low-gap regions, some of which recur across seeds.
The global minimum alone is therefore not sufficient to describe the structure of the problem.
In short: inZOR-ND in this 3D setting reveals a structured family of quasi-degenerate configurations rather than collapse to one basin.