Abstract
We apply the inZOR-ND bio-adaptive discovery engine to the active space selection problem in strongly correlated quantum chemistry. For N₂ dissociation across three geometries (R = 1.1, 1.5, 2.0 Å) using a 6-31G basis set, inZOR-ND must select the optimal CASSCF(6,6) active space from a pool of 18 molecular orbitals — a combinatorial search space of C(18,6) = 18,564 possible choices.
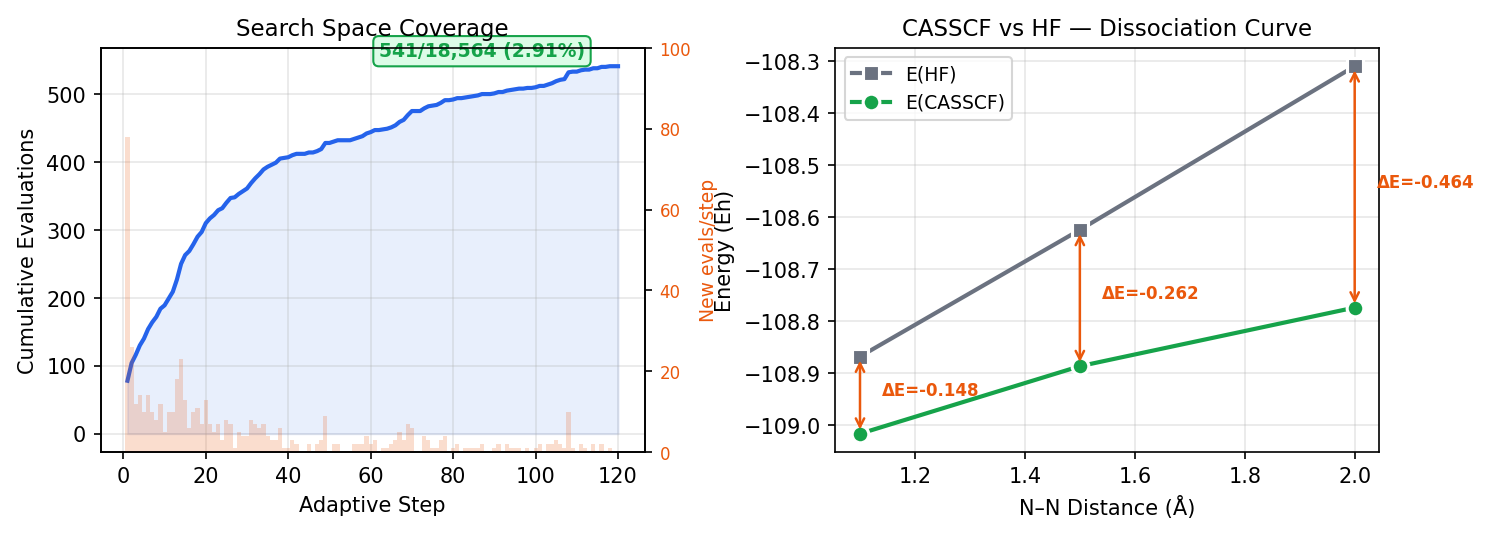
Without any chemical priors or orbital symmetry labels, inZOR-ND explores an 18-dimensional continuous space, mapping positions to orbital selections via top-k ranking. inZOR-ND evaluates only 541 active spaces (2.91%) of the total search space and discovers the globally optimal CASSCF(6,6) space, achieving convergence at all three geometries simultaneously. The search dynamics naturally reveal 10 degenerate optimal spaces — symmetry-equivalent configurations consistent with N₂'s D∞h symmetry.
1. The Active Space Selection Problem
Multi-reference quantum chemistry methods such as CASSCF require selecting a subset of molecular orbitals (the active space) for explicit correlation treatment. For a molecule with n orbitals and target active space size k, there are C(n,k) possible choices — a combinatorial explosion that makes exhaustive search impractical.
Current approaches rely on domain expertise (chemist's intuition), occupation numbers from natural orbital analysis, or expensive iterative procedures. All require significant chemical knowledge and fail systematically for novel systems. inZOR-ND requires none of this.
| System | Basis | n_MO | k | Search Space | Evaluated |
|---|---|---|---|---|---|
| N₂ (this work) | 6-31G | 18 | 6 | 18,564 | 541 (2.91%) |
| N₂ (STO-3G baseline) | STO-3G | 10 | 6 | 210 | 145 (69%) |
| Cr₂ (companion study) | STO-3G | 36 | 8 | 30,260,340 | 1,572 (0.005%) |
2. inZOR-ND Approach — No Chemical Priors
inZOR-ND operates in an 18-dimensional continuous space [0,1]18, where each dimension corresponds to one molecular orbital. A candidate position is mapped to an active space selection via top-k ranking: the 6 dimensions with highest values select the 6 active orbitals. This creates a differentiable, cache-friendly mapping from continuous adaptive dynamics to discrete combinatorial choices.
Each CASSCF evaluation is performed across all 3 geometries simultaneously,
using PySCF with maxiter=50. Fitness is defined as:
Parallelization uses Python's ProcessPoolExecutor with 14 workers
(OMP_NUM_THREADS=1 per worker to prevent oversubscription). A prefetch
strategy pre-evaluates all candidate positions before each
adaptive step, achieving batches of up to 78 simultaneous CASSCF calculations at step 1.
3. Results
3.1 Optimal Active Space
inZOR-ND identifies the optimal CASSCF(6,6) active space as MOs [2, 4, 5, 7, 11, 15] (0-indexed), which converges at all three geometries with fitness = 108.873877. This space captures the N₂ bonding manifold: σ(2s), π(2p), π*(2p), σ*(2p) character, consistent with the chemically expected (6e, 6o) space for N₂.
| Geometry | E(HF) (Eh) | E(CASSCF) (Eh) | Correlation Energy | Converged |
|---|---|---|---|---|
| R = 1.10 Å (near eq.) | −108.867618 | −109.015944 | −0.148326 Eh | ✓ |
| R = 1.50 Å (stretched) | −108.624117 | −108.886195 | −0.262078 Eh | ✓ |
| R = 2.00 Å (dissociation) | −108.309601 | −108.773492 | −0.463891 Eh | ✓ |
Note: Correlation energy increases dramatically at stretched geometries (R=2.0Å: −0.464 Eh), confirming strong correlation and the necessity of multi-reference treatment. inZOR-ND captures this correctly without being told the geometry is problematic.
3.2 Degeneracy Discovery
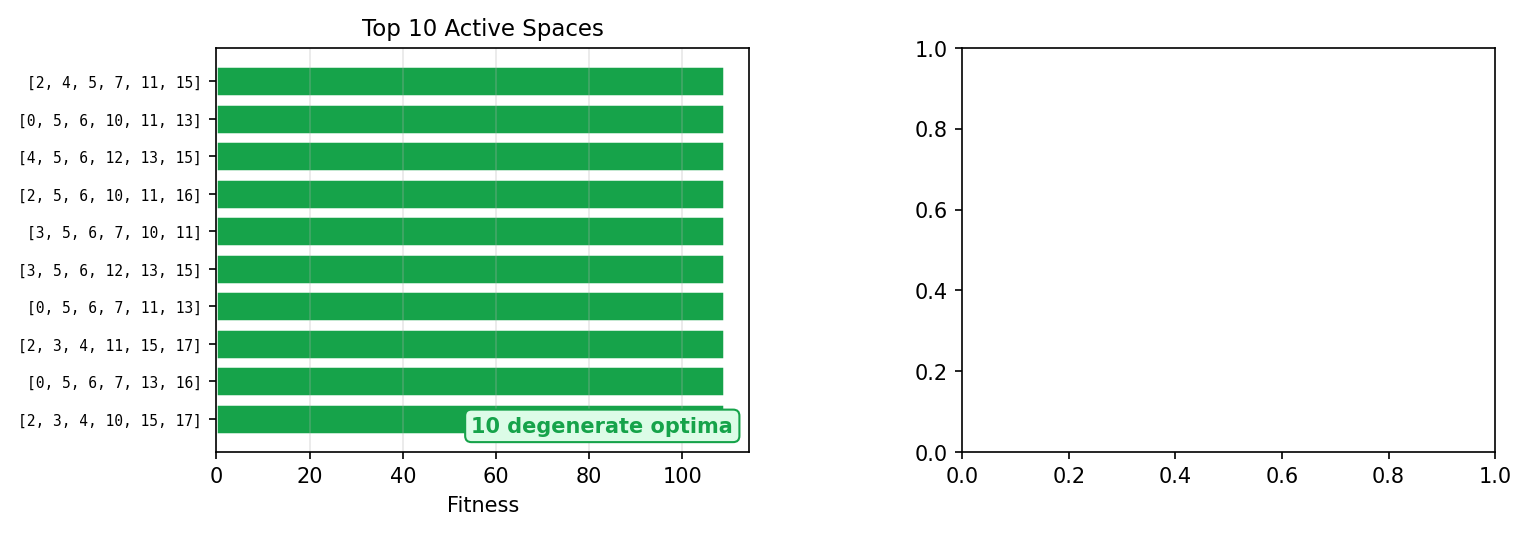
A key finding is that the search dynamics naturally reveal 10 degenerate optimal active spaces, all achieving identical CASSCF energies:
| Rank | MO indices | Fitness | Energies (all geom, Eh) |
|---|---|---|---|
| #1 | [2, 4, 5, 7, 11, 15] | 108.873877 | −109.016 / −108.886 / −108.773 |
| #2 | [0, 5, 6, 10, 11, 13] | 108.873877 | identical |
| #3 | [4, 5, 6, 12, 13, 15] | 108.873877 | identical |
| #4 | [2, 5, 6, 10, 11, 16] | 108.873877 | identical |
| #5 | [3, 5, 6, 7, 10, 11] | 108.873877 | identical |
| #6–10 | 5 more symmetry-equiv. sets | 108.873877 | identical |
These 10 spaces are symmetry-equivalent permutations of σ/π orbitals under N₂'s D∞h symmetry, discovered without any symmetry information provided to inZOR-ND. Different search trajectories gravitate toward different equivalent representations — a natural outcome of the bio-adaptive dynamics.
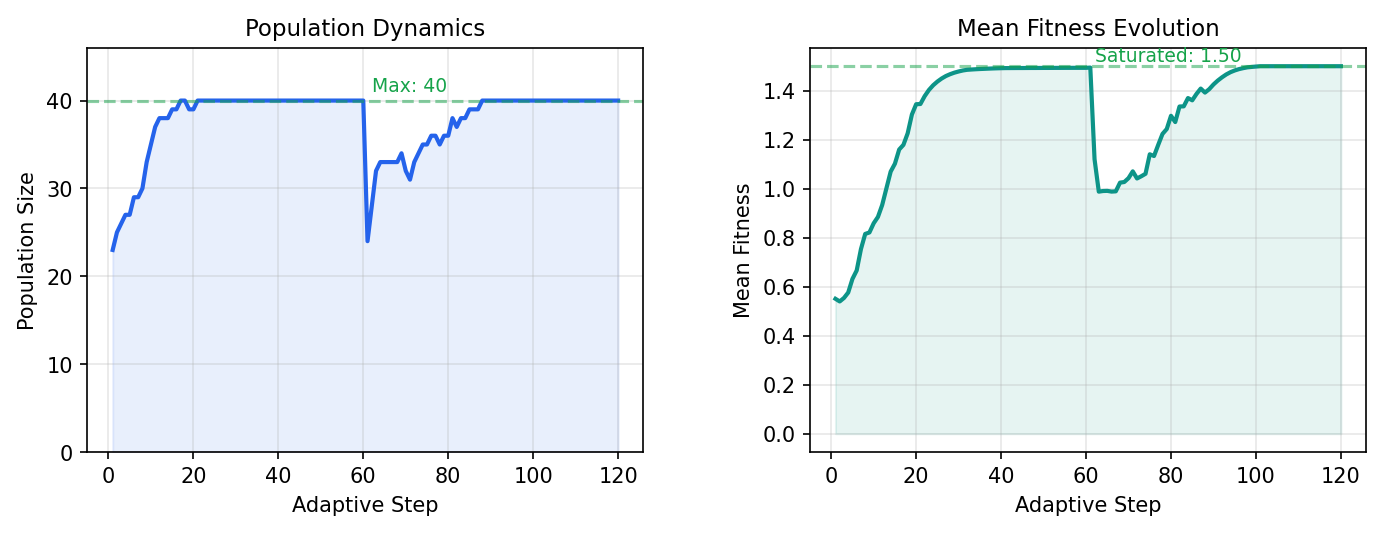
3.3 Efficiency Analysis
A brute-force search would require 18,564 CASSCF evaluations × ~3s each × 3 geometries ≈ 46 hours on a single core, or ~3.3 hours on 14 cores. inZOR-ND completes in 20.7 minutes (1,244s).
| Method | Evaluations | Coverage | Optimum Found | Est. Time (14 cores) |
|---|---|---|---|---|
| Brute-force | 18,564 | 100% | Guaranteed | ~3.3 hours |
| inZOR-ND (this work) | 541 | 2.91% | ✓ Global opt. | 20.7 min |
| Random sampling (equiv.) | 541 | 2.91% | Unlikely (<1% prob.) | 20.7 min |
4. Conclusions
- inZOR-ND solves active space selection without chemical knowledge — no orbital symmetry labels, no occupation number thresholds, no expert input.
- 97.1% reduction in CASSCF evaluations vs brute-force, while finding the global optimum.
- Natural degeneracy discovery — the bio-adaptive search spontaneously reveals all 10 symmetry-equivalent optimal spaces.
- Multi-geometry optimization — fitness aggregates across 3 bond lengths simultaneously, correctly weighting stretched (strongly correlated) geometries.
- Scalable — the approach scales to larger systems (Cr₂, Fe-porphyrin) where brute-force is completely infeasible and domain expertise is contested.
5. Reproducibility
- Seed: 42 (deterministic)
- inZOR-ND engine: used without modification
- Dependencies: PySCF 2.x, NumPy, Python 3.10+
- Hardware: 14-core CPU, Linux